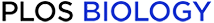Within-host changes across prevalent species of gut bacteria.
(a) Within-host nucleotide differences over 6-month timescales. The blue line shows the distribution of the number of single nucleotide variant (SNV) differences between consecutive quasi-phaseable (QP) time points for different combinations of species, host, and nonoverlapping time interval (if more than two samples are available) for the 45 prevalent species in S20 Fig. The distribution of the number of sites tested in each comparison is shown in S18 Fig. For comparison, the red line shows a matched distribution of the number of SNV differences between each initial time point and a randomly selected Human Microbiome Project host, and the purple line shows the distribution of the number of SNV differences between QP lineages in pairs of adult twins. The shaded regions indicate replacement events (light red, 3% of all within-host comparisons), modification events (light blue, 9% of within-host comparisons), and no detected changes (gray, 88% of within-host comparisons); these ad hoc thresholds were chosen to be conservative in calling modifications. (B) Within-host gene content differences (gains + losses). The blue lines show the distribution of the number of gene content differences within hosts for the samples in (A), with the putative modifications highlighted in light blue, the putative replacements highlighted in light red, and the samples with no SNV changes highlighted in gray. The distribution of the number of genes tested in each comparison is shown in S18 Fig. For comparison, the corresponding between-host and twin distributions are shown as in (A). (C) The total number of nucleotide differences at nondegenerate nonsynonymous sites (1D), 4-fold degenerate synonymous sites (4D), and other sites (2D and 3D) aggregated across the modification events in (A). Sites are stratified based on their prevalence across hosts (S1H Text). For comparison, the gray bars indicate the expected distribution for random de novo mutations (S1H text, part i). (D) The total number of gene loss and gain events among the gene content differences in (B), stratified by the prevalence of the gene across hosts. The de novo expectation for gene losses is computed as in (C); by definition, there are no de novo gene gains.