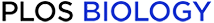Multi-channel models of auxin transport capture observed stem dynamics.
A) The three types of grid structure used to represent tissue organization in the single and multi-channel models. In all three models, cells have dimensions 0.1 mm x 0.01 mm; auxin diffusion rate: D = 16e-3 mm2/min. B) Computer simulations compared to empirical data of the drainage of auxin in an excised stem segment; parameter values are as in C and E, with additional synthesis rates: 3×10−5 auxin units /min-1 (single channel model), 3.8×10−6 auxin units/min-1 (three-channel model). The empirical data are as in Fig 3A. C–E) Computer simulations compared to empirical data of the auxin pulse assay in WT using the single channel model (C), the two-channel model (D), and the three-channel model (E). For the measured data, experiments were as described in Fig 3B and 3C. Error bars show the s.e.m., n = 8 per time point. F) Same simulation as in E, but for the pin1 mutant and later time points. For the measured data, experiments were as described in Fig 3B and 3C, but using pin1-613. Error bars show the s.e.m., n = 8 per time point. Parameter values (mm/min): C. p = 16, q as shown on graph; D. high conductance polar channel: p1 = 6, q1 = 0.1; q12 = 0.; non-polar channel: q3 = 0.01, q32 = 0.01; E. high conductance polar channel: p1 = 2, q1 = 0.2; q12 = 9 10−4; low conductance polar channel: p2 = 0.3, q2 = 0.7, q23 = q22 = q21 = 0.01; non-polar channel: q3 = 0.3, q32 = 2.5 10−4; F. high conductance polar channel: p1 = 0.5, q1 = 0.2; q12 = 9 10−4; low conductance polar: p2 = 0.075, q2 = 0.7, q23 = q22 = q21 = 0.01; non-polar channel: q3 = 0.3, q32 = 2.5 10−4.