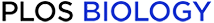Genetic diversity within hosts.
Bacteroides vulgatus is shown as an example in panels A–E; examples for 24 other species are shown in S1 Fig, S2 Fig, and S3 Fig. (A–D) The distribution of major allele frequencies at synonymous sites in the core genome for four different samples, with the median read depth listed above each panel. Major allele frequencies are estimated by max{f,1−f}, where f is the frequency of the base on the reference genome (S1A Text, part iii). To emphasize the distributional patterns, the vertical axis is scaled by an arbitrary normalization constant in each panel, and it is truncated for visibility. The white region denotes the intermediate frequency range used for the polymorphism calculations below. (E) The average fraction of synonymous sites in the core genome with major allele frequencies ≤80% (white region in A–D), for all samples with . Vertical lines denote 95% posterior confidence intervals based on the observed number of counts (S1B Text). The letters indicate the corresponding values for the samples in panels (A–D) for comparison. (F) The distribution of quasi-phaseable (QP) samples among the 35 most prevalent species, arranged by descending prevalence; the distribution across hosts is shown in S7 Fig. For comparison, panels (C) and (D) are classified as QP, while panels (A) and (B) are not.