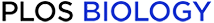Fig 1.tif (459.33 kB)
Gene expression during sea urchin development reflects known phylogenetic relationships between species.
figure
posted on 2016-03-04, 07:57 authored by Jennifer W. Israel, Megan L. Martik, Maria Byrne, Elizabeth C. Raff, Rudolf A. Raff, David R. McClay, Gregory A. Wray(A) Our study includes three sea urchin species: the sister species H. tuberculata (planktotroph) and H. erythrogramma (lecithotroph), which diverged approximately 5 Myr, and an outgroup species L. variegatus (planktotroph), which diverged approximately 35–45 Myr ago [44, 50]. (B) Our developmental time course includes seven stages across each species, from unfertilized eggs to early larvae. (C) Principal component (PC) analysis of gene expression (S1 Data). PC1 explains 36% of the overall variation and clearly separated early (egg through 32-cell stage) from later developmental stages (blastula through early larva), whereas PC2 separated L. variegatus from the two Heliocidaris species, corresponding to their phylogenetic relationships.
History
Usage metrics
Categories
Keywords
life history transitionsSea Urchin Genus Heliocidarisgene expression profilesConserved Gene Regulatory Networkgene expressionGRNoutgroup planktotroph Lytechinus variegatusComparative Developmental TranscriptomicsMajor Life History Switchsea urchin genus Heliocidarisspeciesuse coexpression analysislife history divergencemeasure expression dynamicslecithotroph Heliocidaris erythrogrammalife history conservationsea urchin development
Licence
Exports
RefWorks
BibTeX
Ref. manager
Endnote
DataCite
NLM
DC